Published
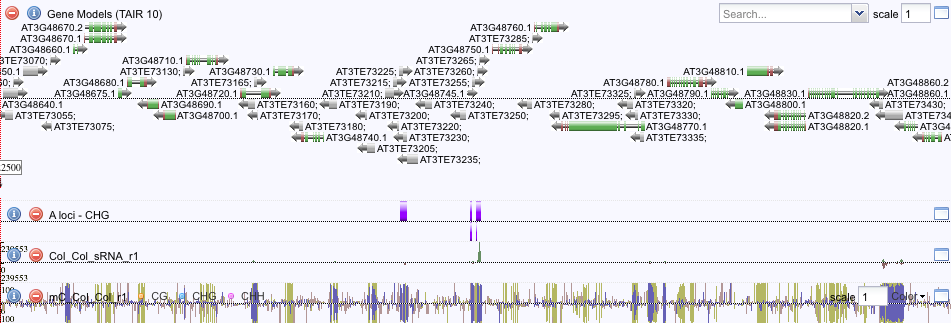
Shoot-root communication of DNA methylationGenome browser of DNA methylation controlled in Arabidopsis thaliana roots controlled by sRNAs from shoots. Experiments conducted by Mat whilst in the Ecker lab in collaboration with Tom Hardcastle and David Baulcombe's lab. Published in PNAS.
|
Unpublished
Transcription factor ChIP-seqGenome browser of example chromatin immunoprecipitation sequencing (ChIP-seq) data mapping binding sites of a transcription factor genome-wide. Generated using the Illumina HiSeq platform. Experiments conducted by Mat whilst in the Ecker lab.
|
Published - Other Labs
The data following were produced by other laboratories. Senior authors and publications are acknowledged in the descriptions. We link to them here because they're particularly interesting!
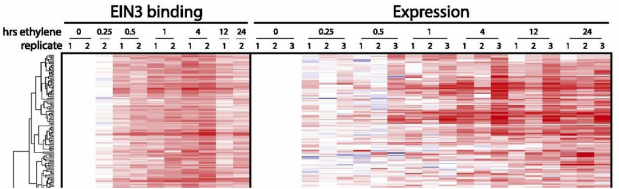
ChIP-seq of EIN3 Genome browser of binding sites for the transcription factor EIN3 mapped in time-series using ChIP-seq. Includes ethylene response time-series transcriptomes. Study published by Kat Chang and the Ecker lab.
|
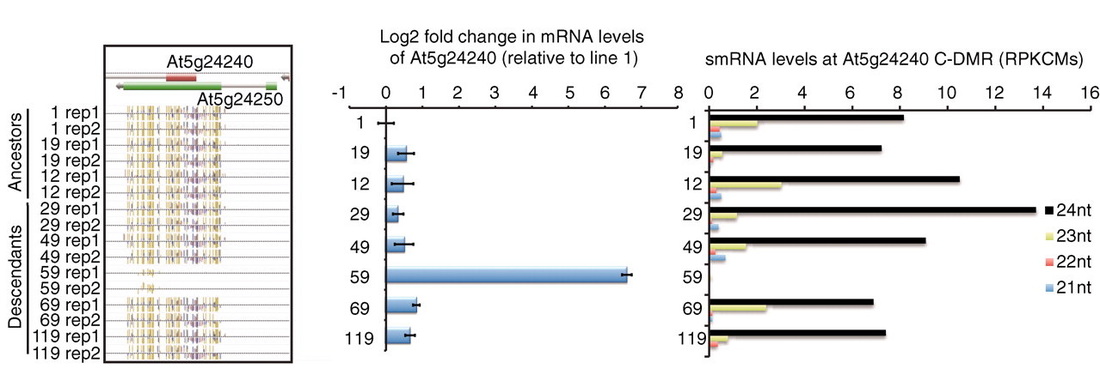
Spontaneous epimutationGenome browser of methylC-seq data that demonstrates spontaneous variation in DNA methylation of the Arabidopsis genome. Published by the Ecker lab and lead by Bob Schmitz.
|
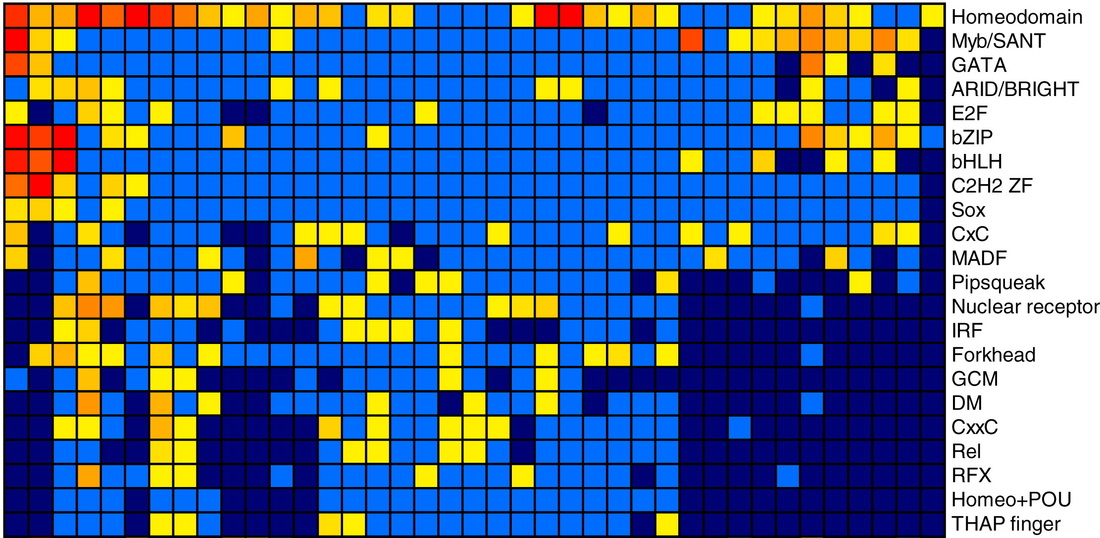
In vitro transcription factor binding assaysDatabase of DNA binding sequence preferences (motifs) for more than 1000 transcription factors from 54 eukaryotic species in vitro. From a published study that used in vitro protein binding microarrays. Carried out by Tim Hughes and Matt Weirauch.
|